| Homeobox
Genes in Development |
4D Microscopy |
Software |
"Virtual Worm"-dbase |
Home
Page |
| Homeobox Genes in Evolution |
Bioinformatics and Modelling |
warthog
& groundhog
Genes |
Teaching |
Homeobox Page |
Homeobox genes are transcription factors that play key roles
in development. The homeodomain is comprised of 3 alpha helices; the
third helix binds to DNA in the major groove (See picture of the Antennapedia Homeodomain). Homeobox
genes fall into many different classes, such as LIM, POU, Prd,
Prd-like, CUT, Antennapedia, etc.
Over the years, we have been, and are working with a series of homeobox
genes that fall into different classes.
For example: ceh-2
(Ems), ceh-8 (prd-like), ceh-14 (LIM), ceh-20 (PBC), ceh-21
(Onecut/cut),
ceh-26 (Pros), ceh-32 (So), ceh-36 (Otd), ceh-37 (Otd), ceh-38
(Onecut/cut),
ceh-43 (Dll), ceh-44 (Cux/cut). We are interested
how these genes control developmental events, in particular during
nervous system development.
Presently, we are studying the developmental role of the Otd homeobox
genes ceh-36, ceh-37, and ttx-1 and their role in
embryogenesis. One of the tools we use is 4D microscopy (see below),
and we have observed early transient expression during embryogenesis
for ceh-36 and ceh-37.
The LIM homeobox gene ceh-14
is another homeobox gene of
particular interest to us. We are currently attempting to identify
target genes of ceh-14 using
gene chip technology and other methods and are planning to study its
regulation during embryogenesis.
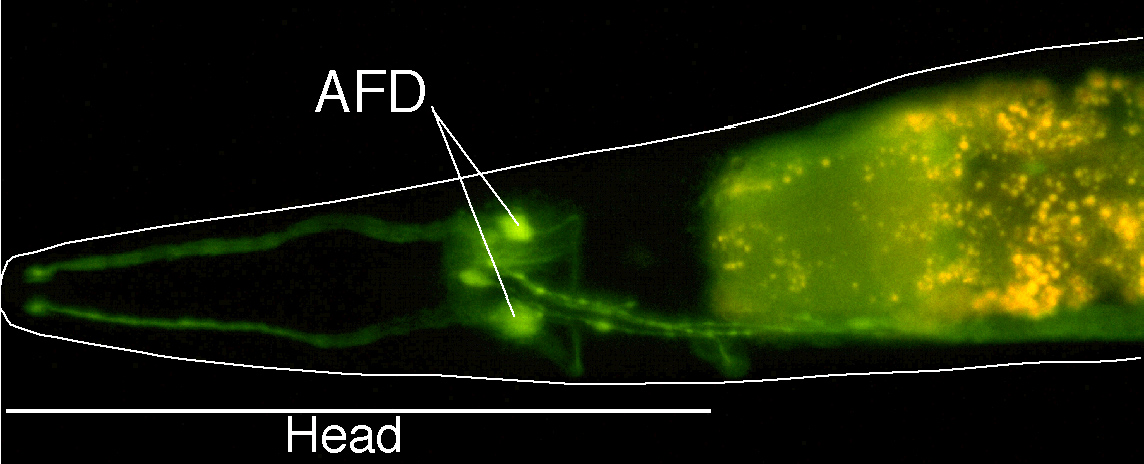
Expression of ceh-14::GFP
in neurons of the head

In collaboration with Genome Canada, British Columbia (Dr. Baillie),
we are also performing an expression survey of those homeobox genes
that have not been studied in detail yet, to identify those, which may
have an interesting role in nervous system development.
4D
Microscopy
 We have developed
software to record not only 4D images during
development, but we can also record live GFP expression patterns of
genes during development. Thus, we can obtain rather precise
information about the spaciotemperal expression pattern of
developmentally important genes. We are applying this technology now to
the analysis of homeobox
gene expression patterns. Some of the software we have developed can be
found on the Software Web
page. To see a GFP movie of a 4D recording, go the the main page.
We have developed
software to record not only 4D images during
development, but we can also record live GFP expression patterns of
genes during development. Thus, we can obtain rather precise
information about the spaciotemperal expression pattern of
developmentally important genes. We are applying this technology now to
the analysis of homeobox
gene expression patterns. Some of the software we have developed can be
found on the Software Web
page. To see a GFP movie of a 4D recording, go the the main page.
 We are interested in
the evolution of homeobox genes to understand
so
as to correlate and correctly identify orthologs when comparing the
functions of these genes across different animal phyla. We have and are
still doing comprehensive analyses of homeobox genes in order to
classify them. We maintain a Web page on Homeobox genes. As part of
this ongoing work, we identified a novel domain in the C-terminus of
HD-ZIP III homeobox genes with putative signalling function (Mukherjee
K, Bürglin TR. (2006) Genome Analysis: MEKHLA, a novel domain with
similarity to PAS domains, is fused to plant homeodomain-leucine zipper
III proteins. Plant Physiol. 140(4):1142-1150).
We are interested in
the evolution of homeobox genes to understand
so
as to correlate and correctly identify orthologs when comparing the
functions of these genes across different animal phyla. We have and are
still doing comprehensive analyses of homeobox genes in order to
classify them. We maintain a Web page on Homeobox genes. As part of
this ongoing work, we identified a novel domain in the C-terminus of
HD-ZIP III homeobox genes with putative signalling function (Mukherjee
K, Bürglin TR. (2006) Genome Analysis: MEKHLA, a novel domain with
similarity to PAS domains, is fused to plant homeodomain-leucine zipper
III proteins. Plant Physiol. 140(4):1142-1150). C. elegans contains a
series of molecules with similarity to the Drosophila signaling
molecule Hedgehog. However, the sequence conservation is restricted to
the C-terminal domain, which has autoproteolytic activity. For a recent
summary of the components of the Hedgehog signaling pathway in C.
elegans see the Wormbook chapter "Homologs
of the Hh signalling network in C.
elegans".
Over 60 open reading frames directly or indirectly related to Hh are
found in C. elegans. We have grouped these genes into several different
families,
termed warthog, groundhog, ground-like, and more recently, quahog.
In a key
paper we intially characterized these genes families:
Aspöck, G., Kagoshima, H., Niklaus, G., and Bürglin, T.R.
(1999). Caenorhabditis
elegans has scores of hedgehog-related
genes:
sequence and expression analysis. Genome Res., 9, 909-923.
Unfortunately, not all supplemental
information for this paper is presented at "Genome Research"journal Web
site,
so here we provide all information: Supplemental
Information.